ENID: The ITEC Endometrial Implants Dataset
ENID (ENdometrial Implants Dataset) comprises 160 images taken from 100+ gynecologic laparoscopy surgeries and is purposefully created to be utilized for a variety of automatic content analysis problems in the context of Endometriosis recognition. ENID is the result of experiments on reorganizing previously published dataset GLENDA in terms of visual similarity.
Description
Endometriosis
Endometriosis is a benign but potentially painful condition among women in child bearing age involving the growth of uterine-like tissue in locations outside of the uterus. Corresponding lesions can be found in various positions and severities, often in multiple instances per patient requiring a physician to determine its extent. This most frequently is accomplished by calculating its magnitude via utilizing the combination of two popular classification systems, the revised American Society for Reproductive Medicine (rASRM) and the European Enzian scores. Endometriosis can not reliably identified by laymen, therefore, the dataset has been created with the help of medical experts in the field of endometriosis treatment.
Purposes
- binary (endometriosis) classification
- detection/localization
Overview
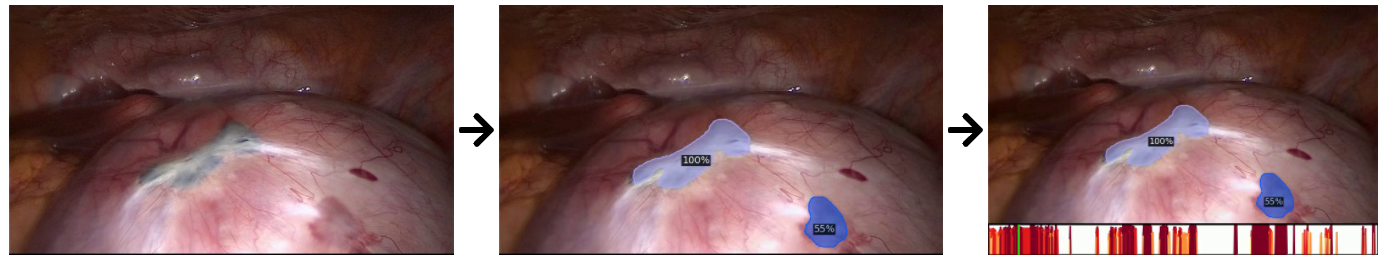
The dataset includes region-based annotations of a specific visual endometriosis manifestation: dark endometrial implants. Annotations are created for single video frames, which can contain multiple annotations. Each single annotation is exported as a pixel-based image mask: the annotated regions are colored, while the background is left black. Additionally a particular dataset augmentation is provided: ground truth annotations are tracked over time using the original videos. The tracking results are acquired via kernelized correlation filters (KCF).
Disclaimer
The dataset is exclusively provided for scientific research purposes and as such cannot be used commercially or for any other purpose. If any other purpose is intended, you may directly contact the originator of the videos, Prof. Dr. Jörg Keckstein.
In addition, reference must be made to the following publication when this dataset is used in any academic and research reports:
A. Leibetseder, K. Schoeffmann, J. Keckstein and S. Keckstein. 2021. Endometriosis Detection and Localization in Laparoscopic Gynecology. Accepted: TBA.
BibTeX:
@article{DBLP:journals/mta/LeibetsederSKK22,
author = {Andreas Leibetseder and
Klaus Schoeffmann and
J{\"{o}}rg Keckstein and
Simon Keckstein},
title = {Endometriosis detection and localization in laparoscopic gynecology},
journal = {Multim. Tools Appl.},
volume = {81},
number = {5},
pages = {6191--6215},
year = {2022},
url = {https://doi.org/10.1007/s11042-021-11730-1},
doi = {10.1007/s11042-021-11730-1},
timestamp = {Thu, 03 Mar 2022 09:23:23 +0100},
biburl = {https://dblp.org/rec/journals/mta/LeibetsederSKK22.bib},
bibsource = {dblp computer science bibliography, https://dblp.org}
}
ENID is licensed under Creative Commons Attribution-NonCommercial 4.0 International (CC BY-NC 4.0,  ) and is created as well as maintained by Distributed Multimedia Systems Group of the Institute of Information Technology (ITEC) at Alpen-Adria Universität in Klagenfurt, Austria.
) and is created as well as maintained by Distributed Multimedia Systems Group of the Institute of Information Technology (ITEC) at Alpen-Adria Universität in Klagenfurt, Austria.
This license allows users of this dataset to copy, distribute and transmit the work under the following conditions:
- Attribution: You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
- Non-Commercial: You may not use the material for commercial purposes.
Dataset
Following table shows dataset statistics and provides preview/download links. Please note that due to the large number of images in the tracked dataset, it can only be previewed once downloaded.
IMPORTANT: By downloading, you agree to and adhere to above conditions.
| dataset | cases | frames | annotations | size | preview | download |
|---|---|---|---|---|---|---|
| ENID | 108 | 160 | 358 | 12MB | ||
| tracked ENID | 108 | 12 533 | 18 275 | 973MB | - |
Splits
Below both of accompanying research's splits for untracked ENID can be downloaded, i.e. 60/20/20 and 80/10/10. For split preview to work, please extract either of archives into the main dataset folder.
IMPORTANT: By downloading, you agree to and adhere to above conditions.
Models
Below table lists the performance of the two best augmented pre-trained Mask R-CNN models of both above splits in comparison to the respective best raw (unaugmented) model. Additionally, a comparison view per split gives insights over the models' qualitative performance. All models have been trained using Detectron2 and the initial model weights are set according to selected models from the Detectron2 Model Zoo (referenced by 'base model' column).
IMPORTANT: By downloading, you agree to and adhere to above conditions.
Demo
A demo application has been published at CBMI 2021 and can be found at https://github.com/amplejoe/EndometriosisSegmentationTool.